Software at CIS : lddmm-landmark : Validation
manual | faq | input formats | validation | credits | changelog | feedback
Content:
Overview
The LDDMM Validation section provides input data, processing and visualization examples for LDDMM to ensure correctness of the resultant data. These examples are useful tests when LDDMM is run on new environments or platforms. Examples are displayed with available visualization software. A sample lddmm-landmark command is posted with each example (or type lddmm-landmark --help for more lddmm-landmark usage information).
X to H Test
Input: input.tar.gz
Output: output.tar.gz
DTIStudio Files: dtistudio.tar.gz
Command: lddmm-landmark -s 7.5 -j -m -c -w LDDMMMatrix.hdr -p ThreeLayerX.txt ThreeLayerX.txt ThreeLayerH.txt
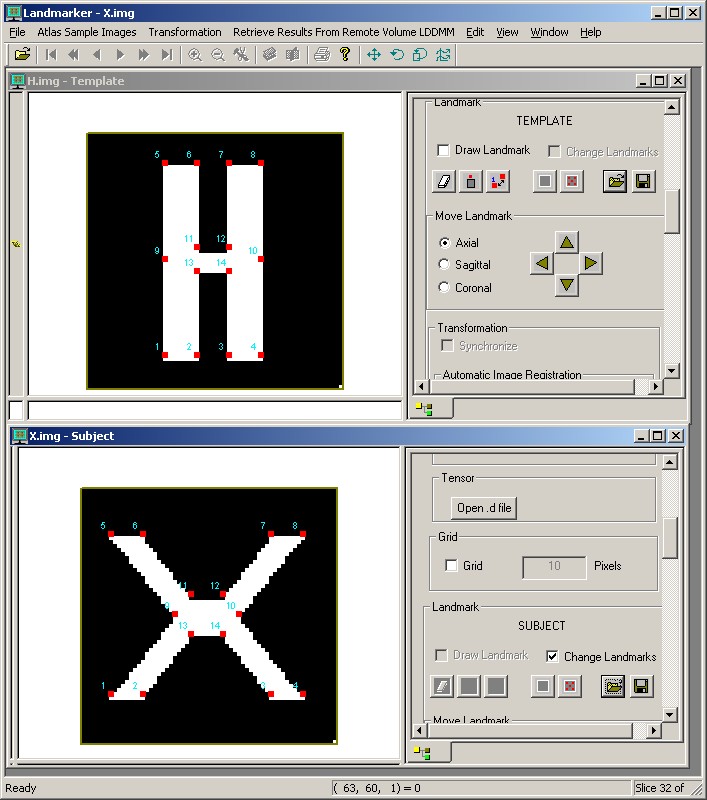
This test matches landmarks on an "X" to landmarks on an "H" using lddmm-landmark on a Unix platform. The original landmarks are then flowed according to the lddmm results, and a volume transformation (*.xfm file) is created.
This matching can similarly be performed in Landmarker. The H image and
the associated landmarks are loaded as the template, and the X is loaded as
the subject. The user selects
Transformation->Landmark Based LDDMM
from the menu,
then performs a 2D transformation with 3 slices and a 1 pixel interval,
and a sigma of 7.5.
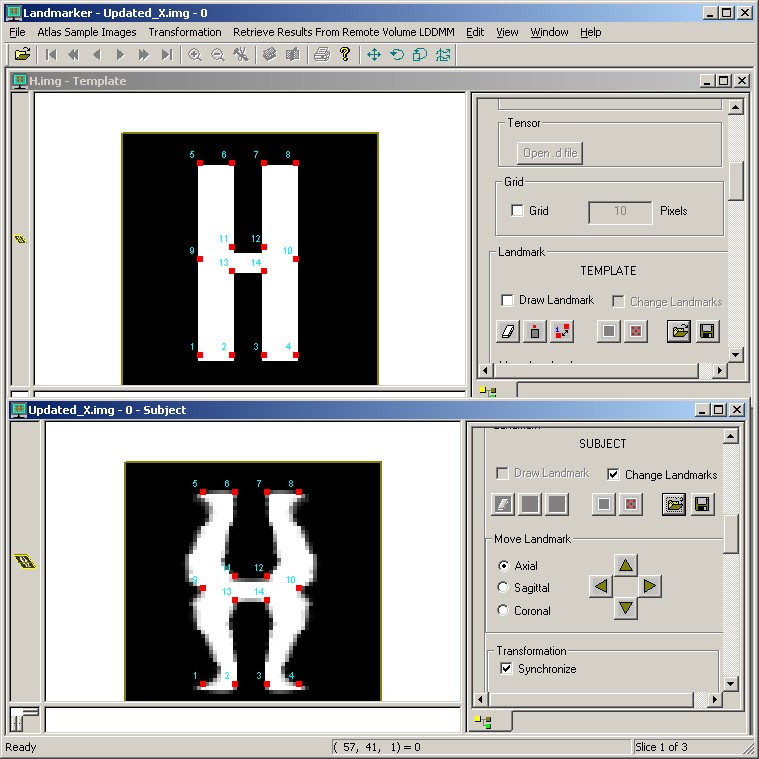
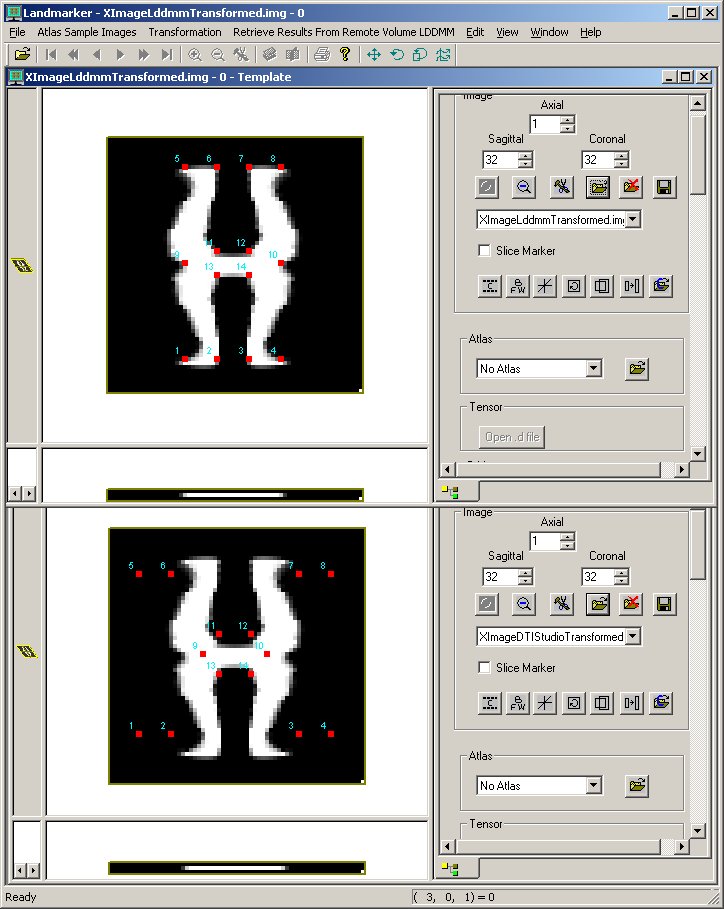
The following figures show the results of matching as displayed in Landmarker. The first figure is the input data before matching. The second is after the X has been deformed by lddmm-landmark matching in Landmarker. The third shows the X image after the transform computed on the Linux platform is applied.
Sphere to Ellipse Test
Input: input.tar.gz
Output: output.tar.gz
Image Files: images.tar.gz
Command: lddmm-landmark -s 10.0 -j -m -c -A sphere.vti -p largeSphere.lmk templateSphere.lmk targetEllipse.lmk
This test matches landmarks on a sphere to corresponding landmarks on an ellipse using lddmm-landmark on a Unix platform.
After matching, first a larger file of points randomly uniformly distributed on the sphere are flowed according to the lddmm-landmark results. The sphere is deformed into the elliptical shaped object. Second, image data that represents a sphere is deformed. The sphere is also deformed to the shape of an ellipse.
The following figures show first the results of matching as displayed in CAWorks. Pictured first are the spherical points (blue) input into lddmm-landmark, the elliptical points (yellow) they are matched with.
| Template Sphere (blue) and Target Ellipse (yellow) |
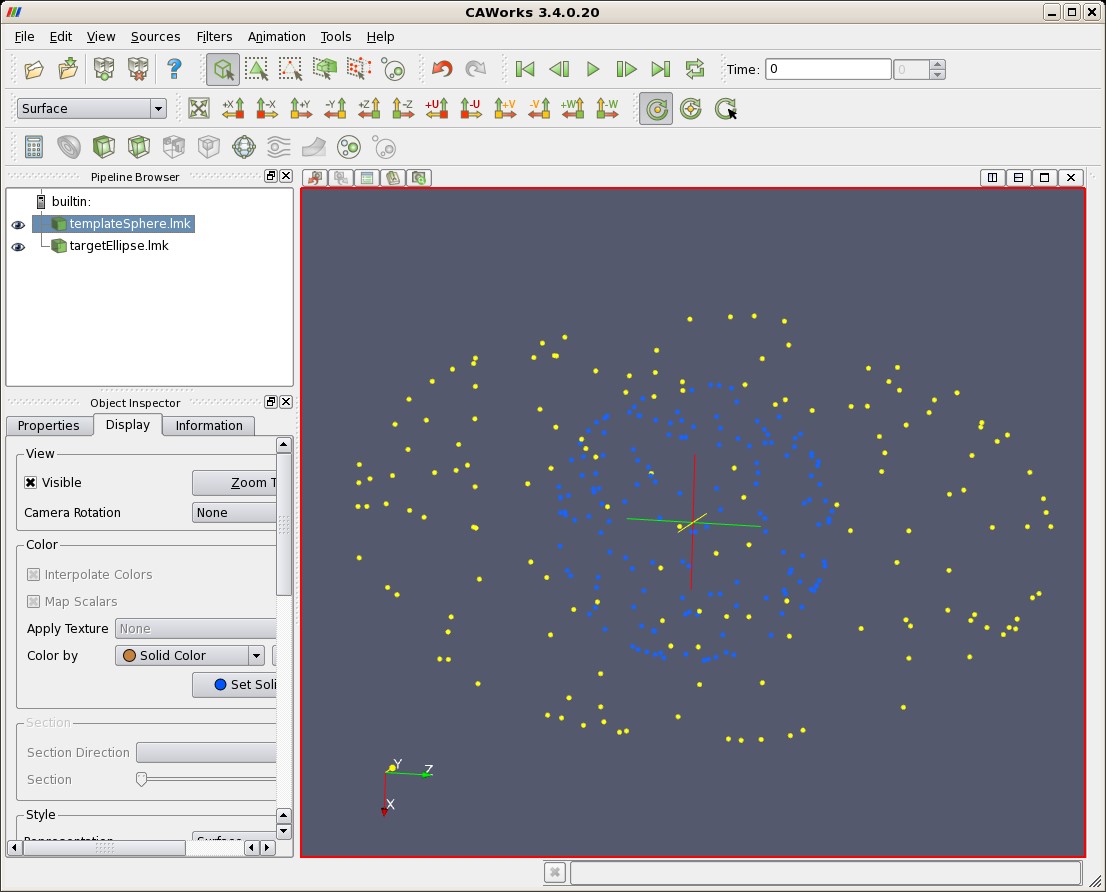
|
| Deformed Template Sphere (blue) and Target Ellipse (yellow) |
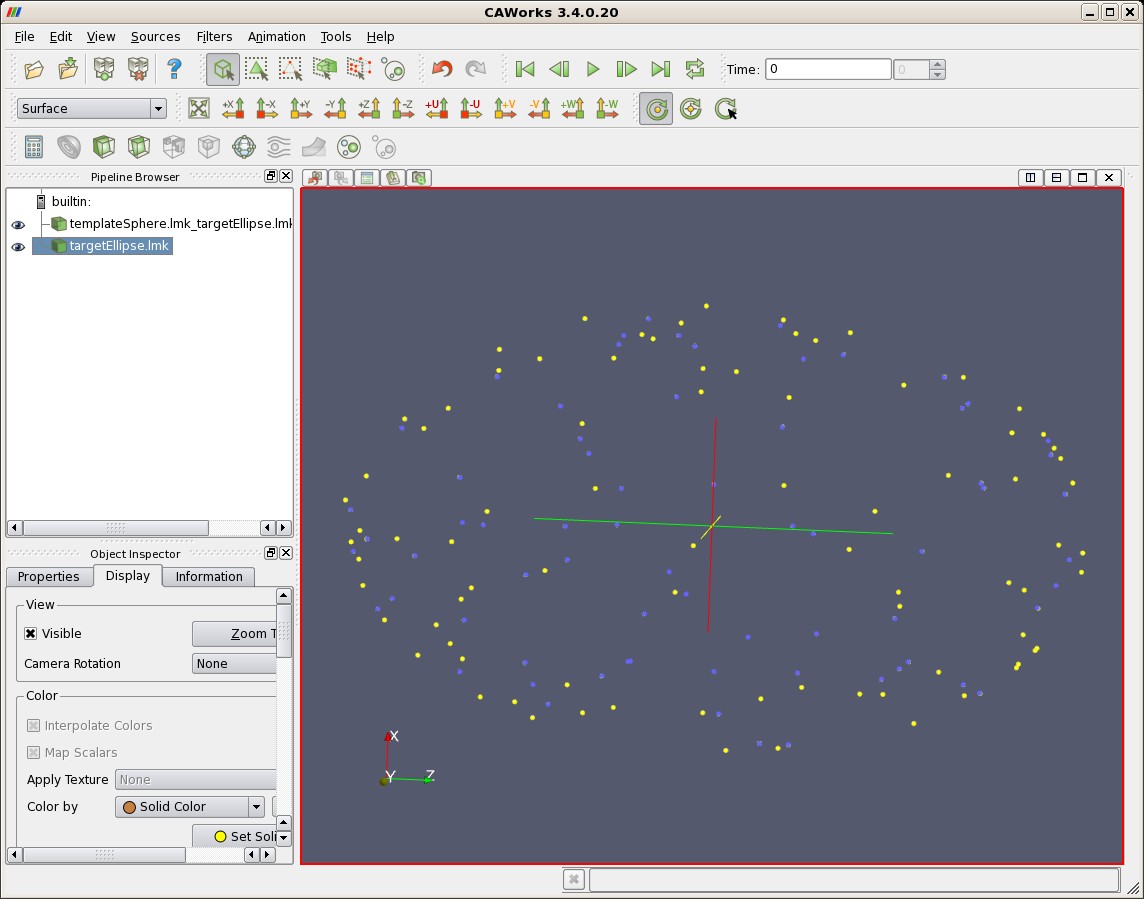
|
| Landmarks Uniformly Distributed on Sphere (red) and Target Ellipse (white) |
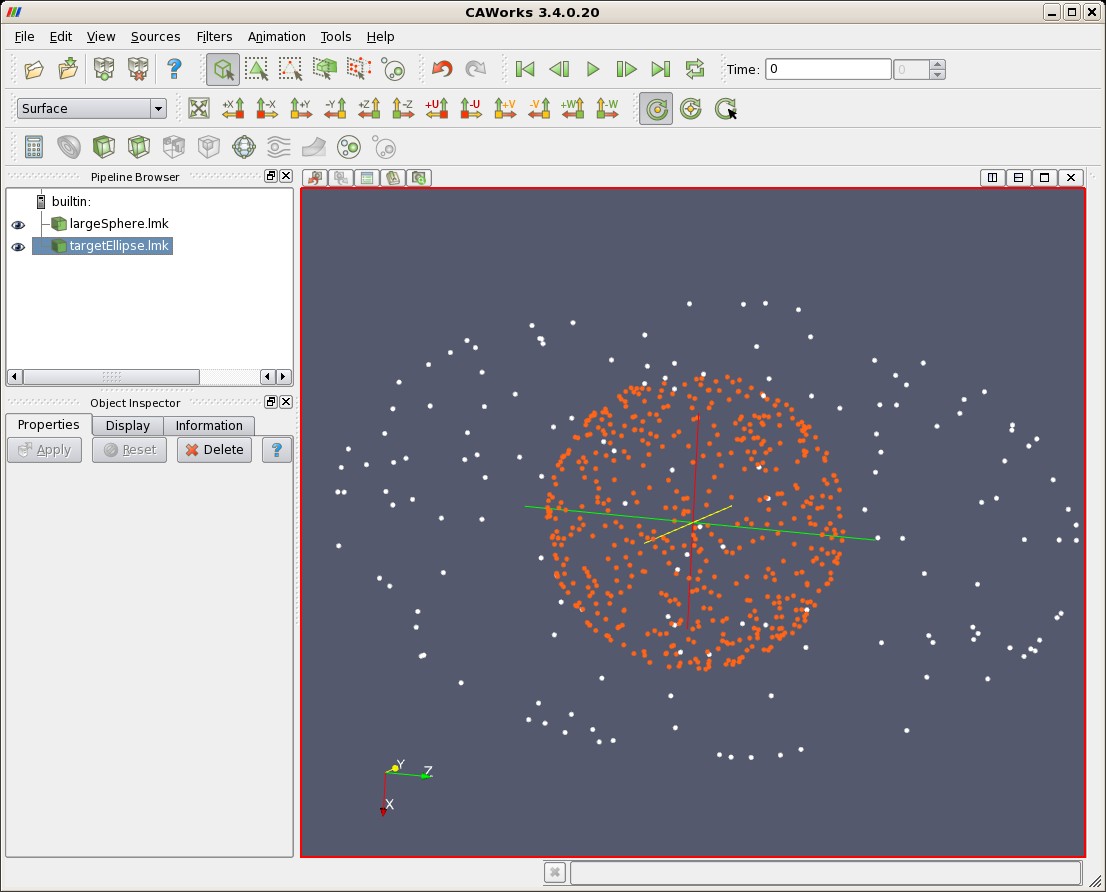
|
| Landmarks (red) Deformed According to Matching Results, and Target Ellipse (white) |
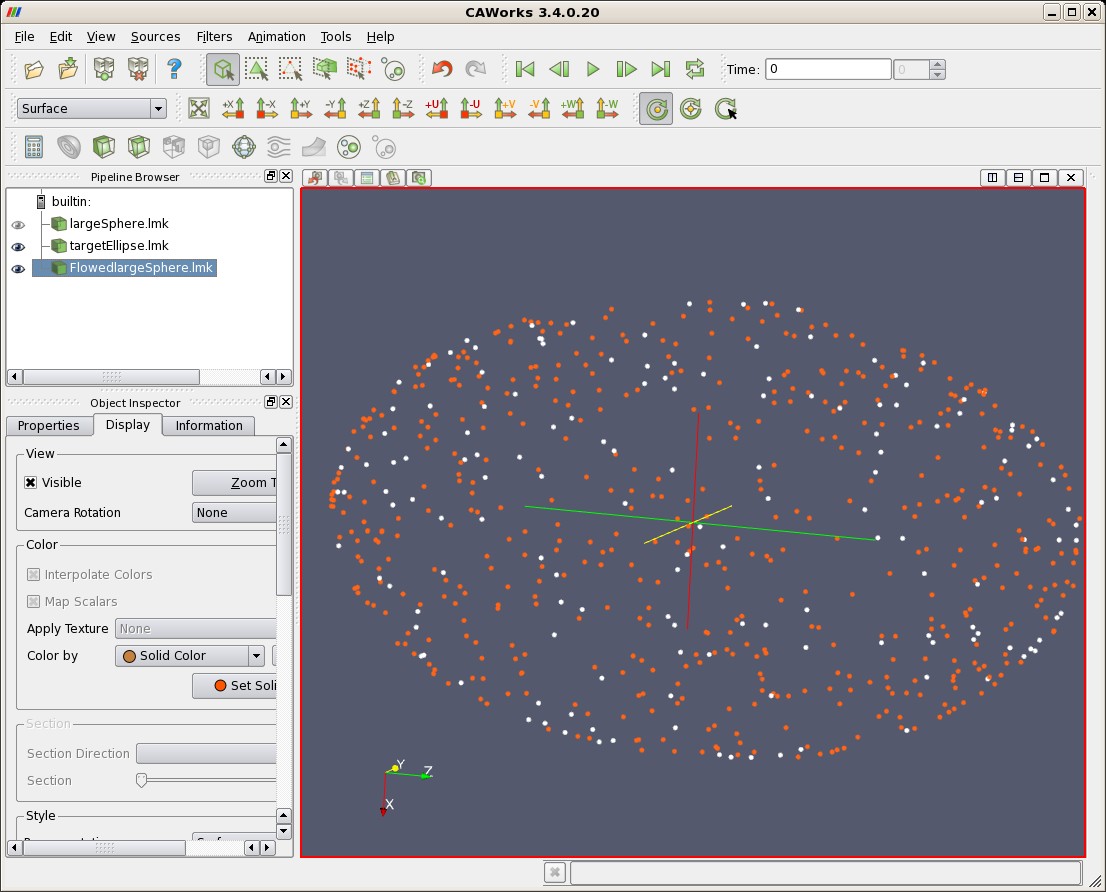
|
| Sphere (gray) Deformed According to Matching Results, and Target Ellipse (yellow) |
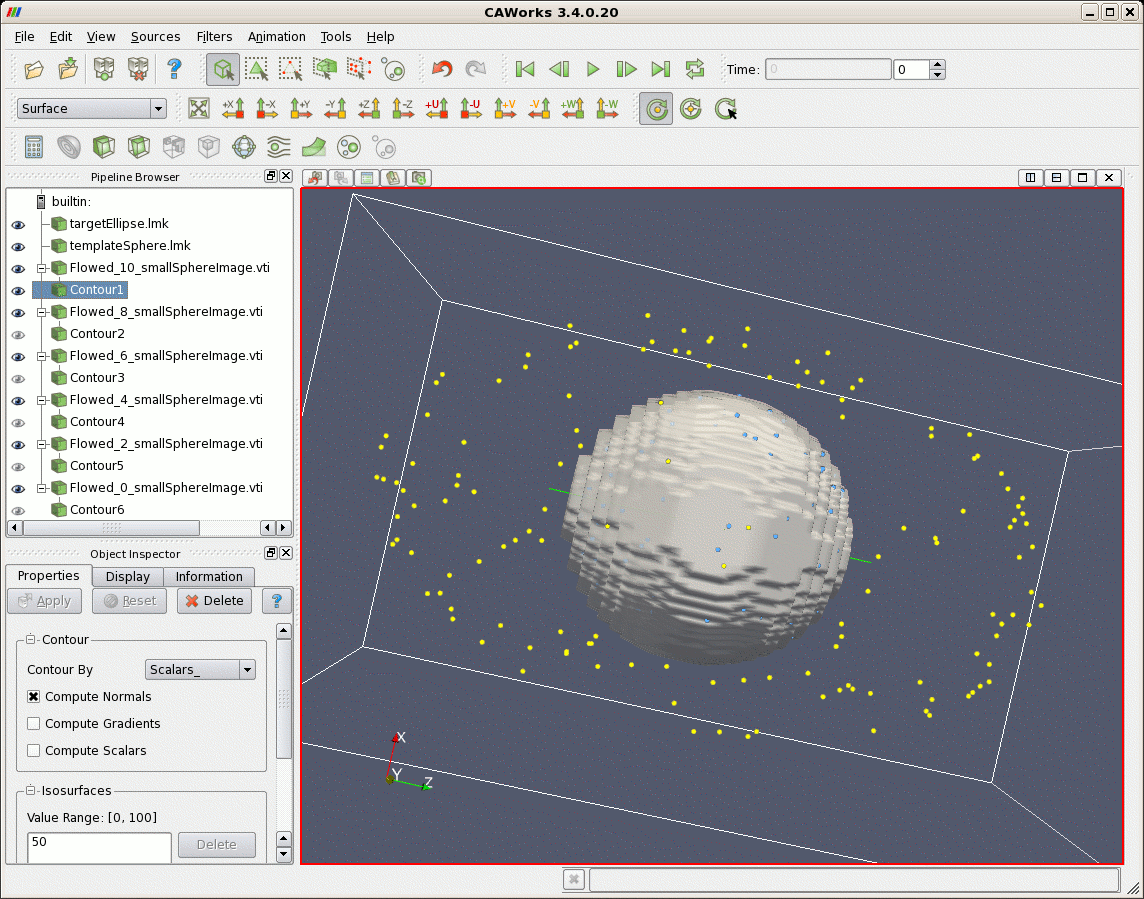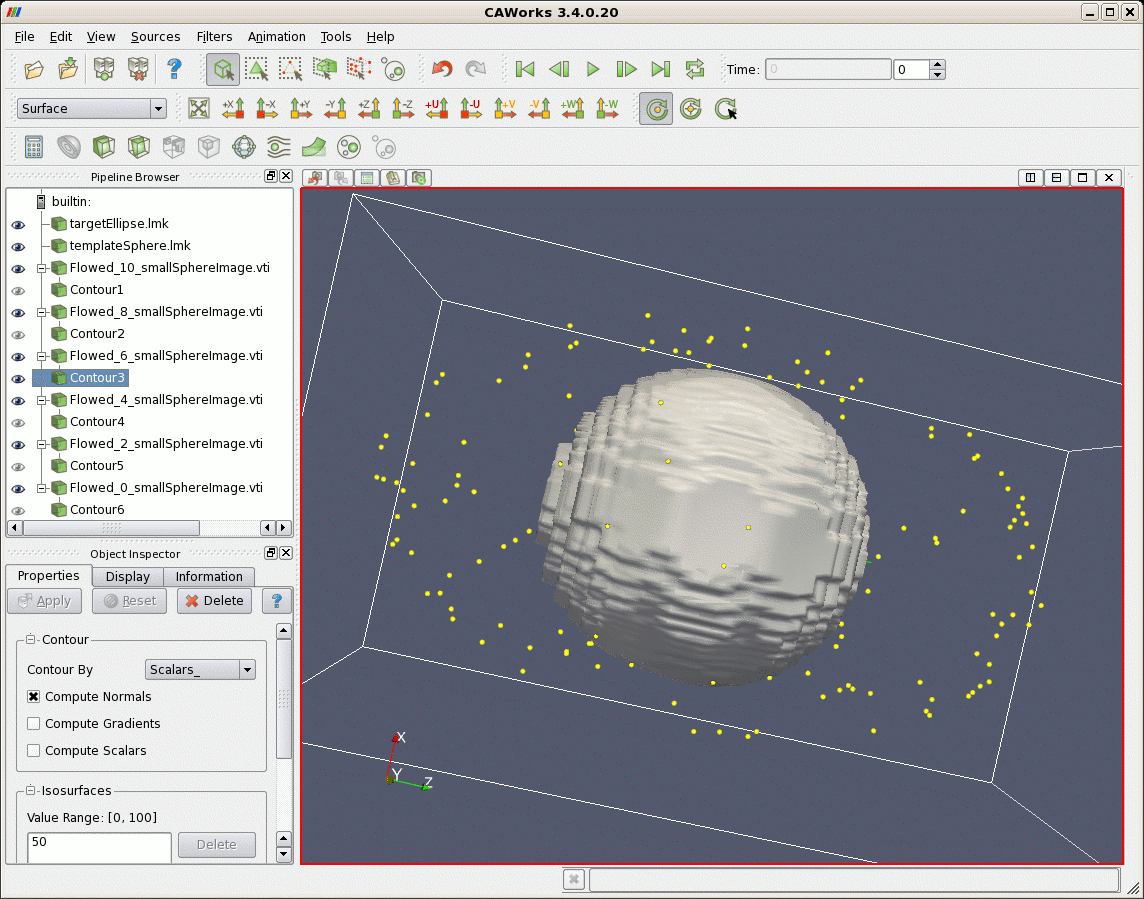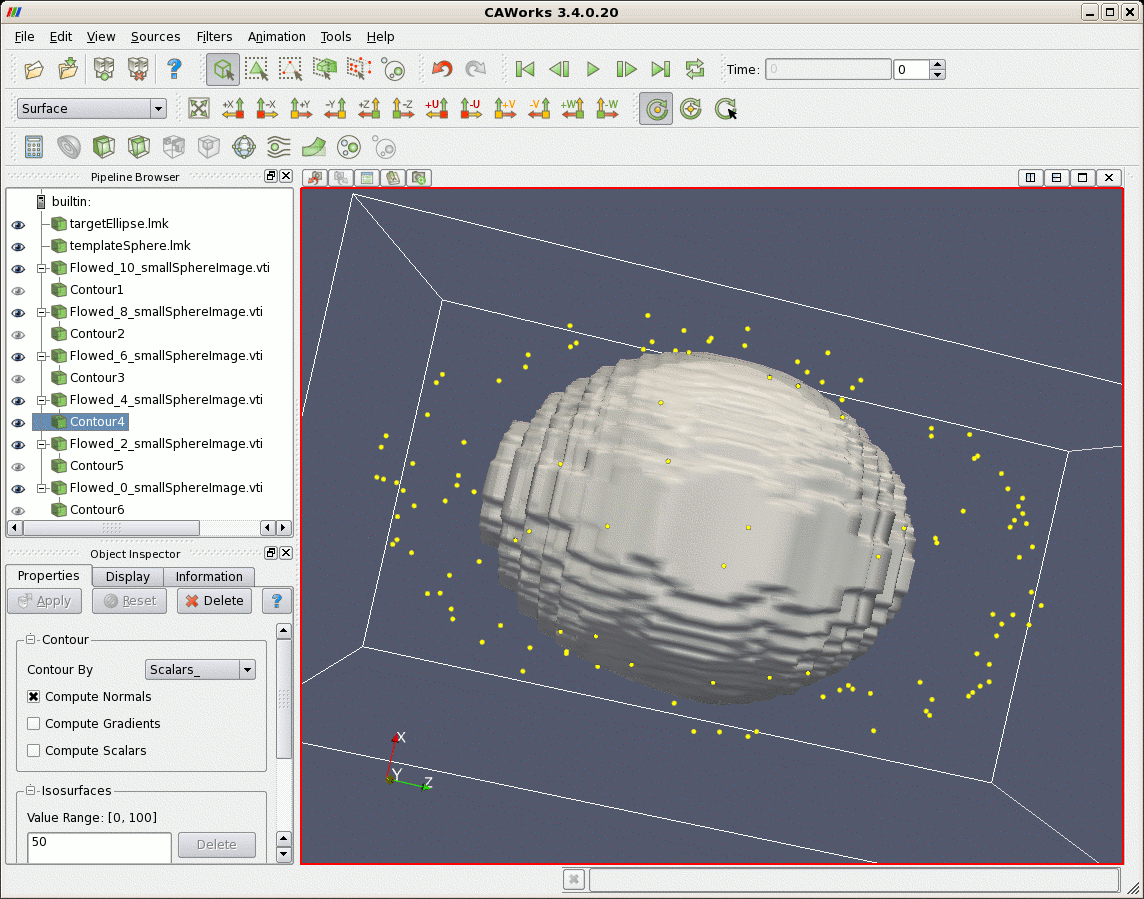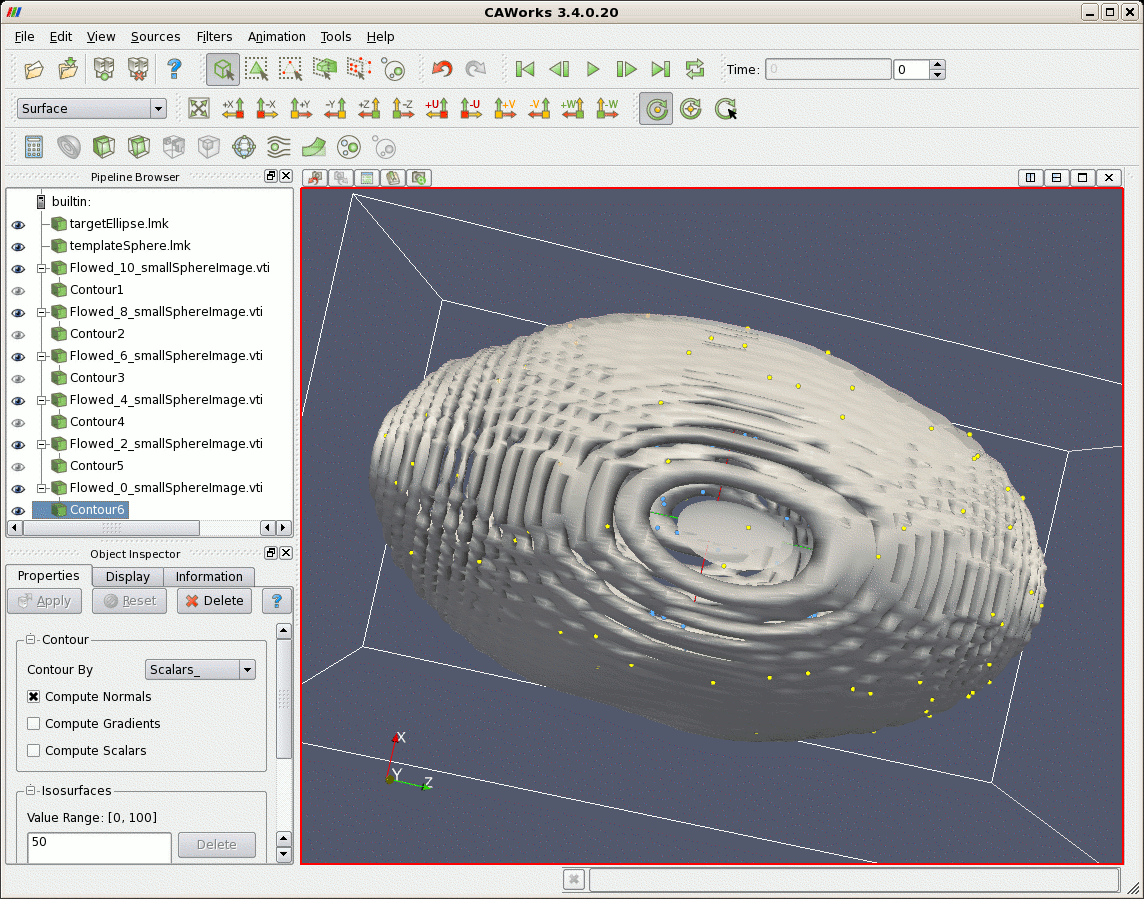
|
Monkey Brain Test
Input: input.tar.gz
Output: output.tar.gz
DTIStudio Files: dtistudio.tar.gz
Command: lddmm-landmark -s 7.5 -j -m -c -w LDDMMMatrix.hdr monkey1.lmk monkey2.lmk
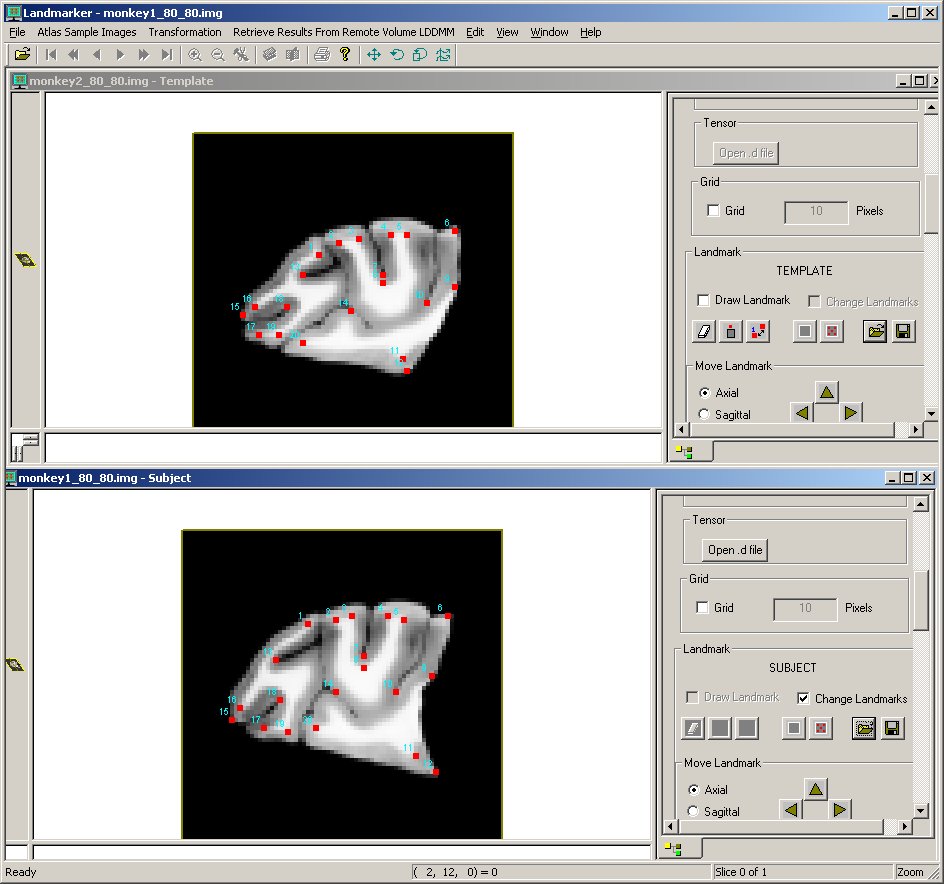
This test matches landmarks on two monkey brains using lddmm-landmark on a Unix platform. A volume transformation (monkey1.lmk_monkey2.lmk.xfm) is created during the matching.
After matching, the monkey brain Analyze image files are loaded into DTIStudio Landmarker (monkey2 = template, monkey1 = subject). The VolumeTransformation file is loaded onto the subject file, and the target landmarks superimposed on the image.
The following figures show the results of matching as displayed in DTIStudio Landmarker. Pictured here are the image and landmark data before and after the volume transformation is applied.
Hippocampus Apply Deformation Test
Input: input.tar.gz
Output: output.tar.gz
Command: lddmm-landmark -A y7_L_handseg.img -B -C -s 5 -t 20 -c -l 50 -E LDDMMMatrix.hdr -F LDDMMMatrix.hdr y7_1.Lf0_38.lmk y1_1.Lf0_38.lmk
This test matches landmarks placed on a hippocampus using lddmm-landmark on a Unix x86_64 platform. The deformation results are applied to the Analyze Volume y7_L_handseg.img, which is a flipped binary segmentation of the template volume. The -B option flips the volume into proper xyz space and -C requests Nearest Neighbor interpolation. The -E option requests an Hmap of the deformation in the volume space specified in the file LDDMMMatrix.hdr. The -F does the same, except that the vector filed values are relative to the volume's voxel positions (vs. absolute xyz positions).
The following figures show the results of matching as displayed in CAWorks. The first figure is an isosurface of the template binary segmentation and a slice of the differential vector field. The second contains isosurfaces of the deformed template and target binary segmentations.
| Template Binary Segmentation Isosurface and Slice of Differential Hmap |
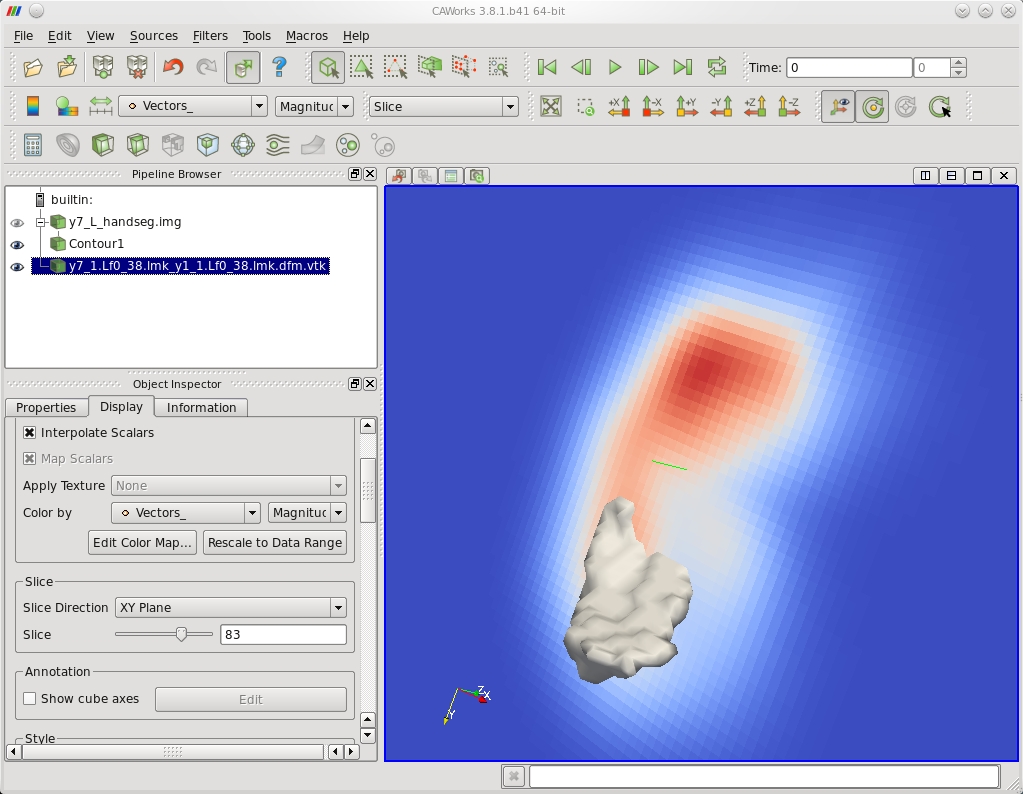
|
| Template Hippocampus Binary Segmentation y7 Matched to Target Hippocampus y1 |
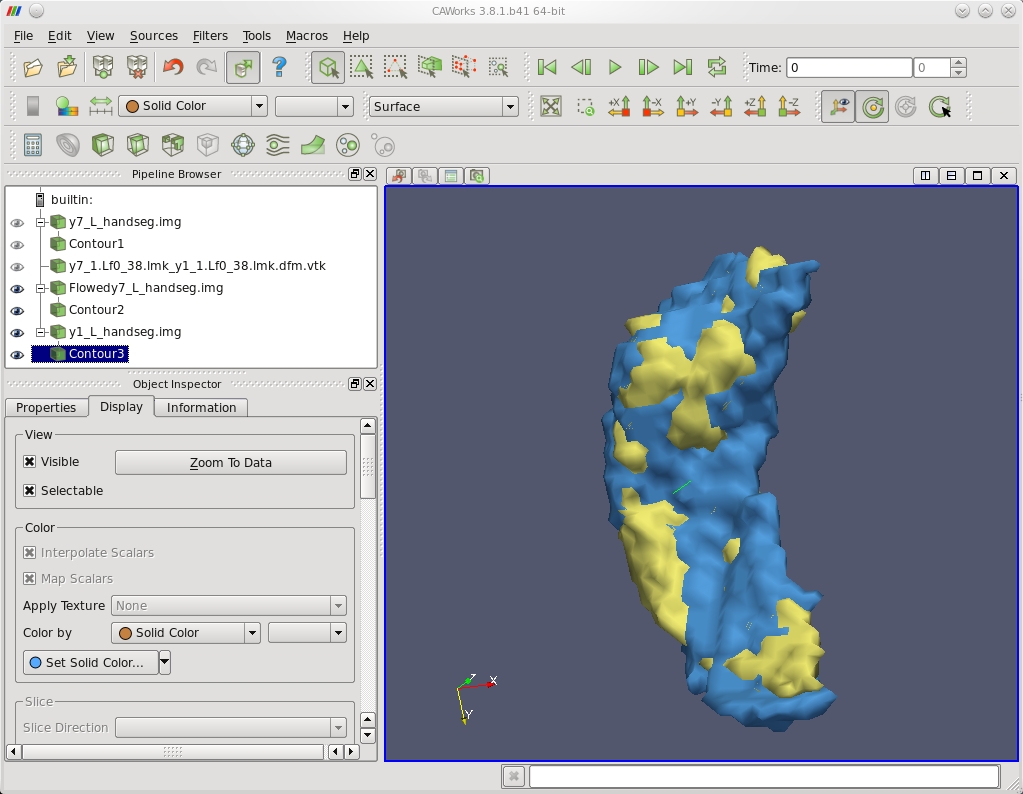
|
Last Modified: Monday, 14th March, 2011 @ 05:10pm